Dynamic self-assembly of molecular capsules via solvent polarity controlled reversible binding of nitrate anions with C 3 symmetric tripodal receptors ... - Chemical Communications (RSC Publishing) DOI:10.1039/C1CC12757H

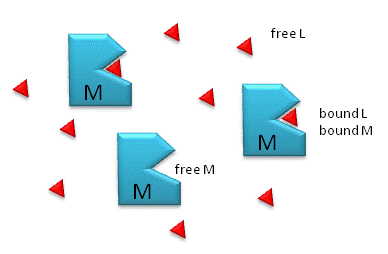
Binding mechanism of reversible, slowly/partially irreversible, and... | Download Scientific Diagram
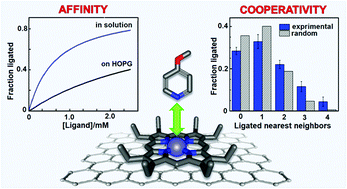
Quantifying reversible nitrogenous ligand binding to Co(ii) porphyrin receptors at the solution/solid interface and in solution - Physical Chemistry Chemical Physics (RSC Publishing)

Distinguishing the optimal binding mechanism through reversible and irreversible inhibition analysis of HSP72 protein in cancer therapy - ScienceDirect
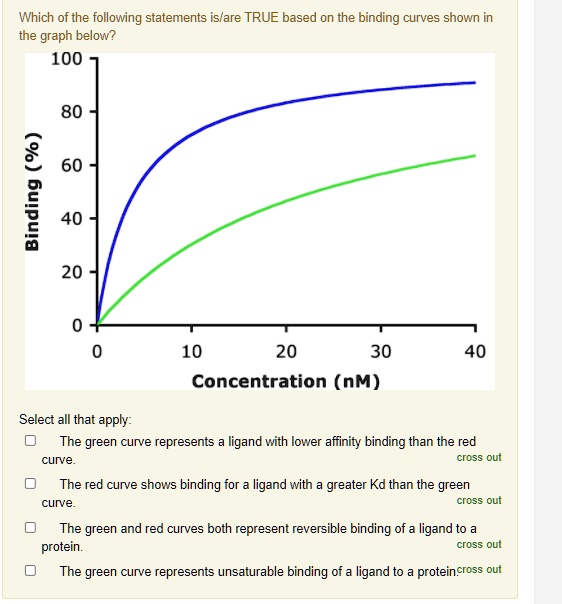
SOLVED: Which of the following statements islare TRUE based on the binding curves shown in the graph below? 100 80 2 60 I 40 20 10 20 30 Concentration (nM) 40 Select

Reversible CO Binding Enables Tunable CO/H2 and CO/N2 Separations in Metal-Organic Frameworks with Exposed Divalent Metal Cations | Berend Smit

Simplified free energy profile of two-state binding for a reversible... | Download Scientific Diagram
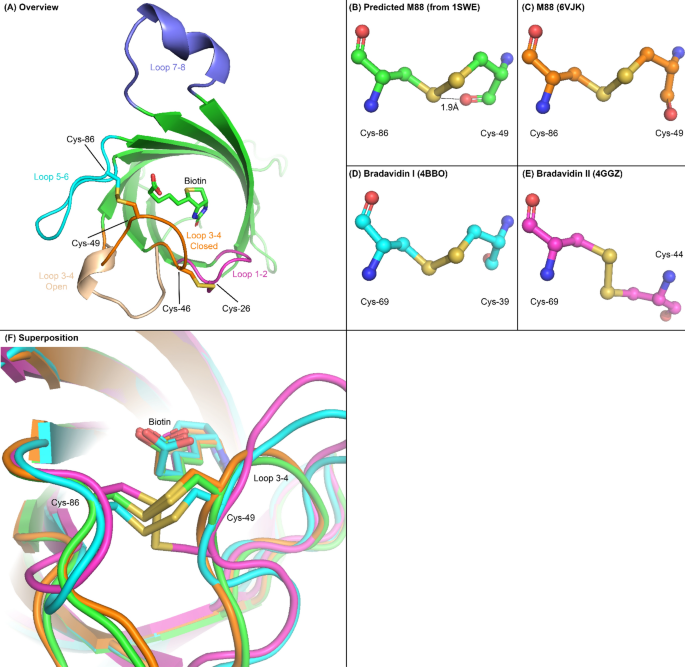
Engineering a disulfide-gated switch in streptavidin enables reversible binding without sacrificing binding affinity | Scientific Reports

Equilibrium and Kinetic Measurements Reveal Rapidly Reversible Binding of Ras to Raf (∗) - Journal of Biological Chemistry

Cell‐Active, Reversible, and Irreversible Covalent Inhibitors That Selectively Target the Catalytic Lysine of BCR‐ABL Kinase - Chen - 2022 - Angewandte Chemie International Edition - Wiley Online Library

Novel Reversible-Binding PET Ligands for Imaging Monoacylglycerol Lipase Based on the Piperazinyl Azetidine Scaffold | Journal of Medicinal Chemistry
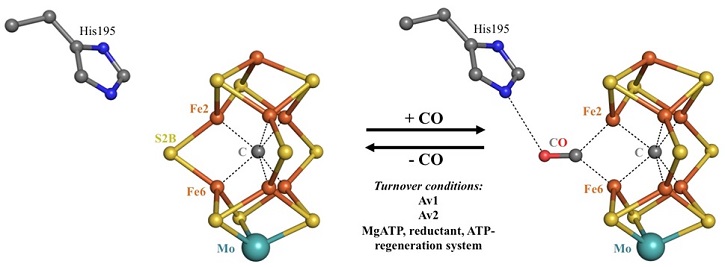
Reversible CO-binding to the Active Site of Nitrogenase | Stanford Synchrotron Radiation Lightsource

Recombinant Antibody Engineering Enables Reversible Binding for Continuous Protein Biosensing | ACS Sensors